Hierarchical Segmentation
During resolution reduction by diffusion, local signal variations smooth out as ''time'' t proceeds. Extrema disappear one after the other (only rarely a new on in created, see [6]), and finally only the strongest signal variations survive. Thus a hierarchical ordering of extrema is induced by the diffusion process.
The images f(x,y;t) obtained from the original
data f(x,y;t) obey further a concept of causality
[3]: every value ![]() found at a specific scale
found at a specific scale ![]() can be traced back to the original resolution t=0. The contour sheets
can be traced back to the original resolution t=0. The contour sheets
![]() are, except for a finite number of critical values
are, except for a finite number of critical values ![]() , two-dimensional submanifolds of
, two-dimensional submanifolds of ![]() . This follows directly from Sard's theorem. At the critical values
. This follows directly from Sard's theorem. At the critical values ![]() , the topology of the contour sheets changes. These events correspond to
the aforementioned disappearance or appearance of local extrema.
, the topology of the contour sheets changes. These events correspond to
the aforementioned disappearance or appearance of local extrema.
From Thom's theory it follows that every such event has qualitatively the same generic form [3]. Since f(x,y;t) is a one-parameter family of functions in t, the only possible way to change is given by the so-called fold catastrophe, generically described through the function
![]()
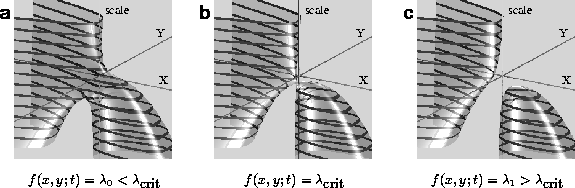
In figure 2 the contour sheets
of this function are displayed near a critical value ![]() . For
. For ![]() , we see a branching of the contour-sheet as one descends towards finer
resolutions (figure 2.a).
, we see a branching of the contour-sheet as one descends towards finer
resolutions (figure 2.a).
Figure 2: The change of the topology of contour sheets near a critical
value ![]() . Connected at
. Connected at ![]() (a), the sheet splits into two disjunct sets for
(a), the sheet splits into two disjunct sets for ![]() (c).
(c).
For ![]() (figure 2.c), the manifold separates
into two disjunct pieces, one continuing towards coarser resolutions, and
the other one descending in scale space towards finer resolutions. Clearly,
the surface spawned from the main surface is orientable, and encircles
an area at the finest resolution which is topological equivalent to a disk
(or a union of discs if it branches again).
(figure 2.c), the manifold separates
into two disjunct pieces, one continuing towards coarser resolutions, and
the other one descending in scale space towards finer resolutions. Clearly,
the surface spawned from the main surface is orientable, and encircles
an area at the finest resolution which is topological equivalent to a disk
(or a union of discs if it branches again).
Between critical values, all surfaces are similar and can be continuously
deformed into each other. We thus have an onion-like, hierarchical structure
of contour sheets, changing topology only at a few critical values ![]() , corresponding to the disappearance of local extrema present in the input
data.
, corresponding to the disappearance of local extrema present in the input
data.
Several schemes have been proposed for exploiting the structure of scale
space [3, 6,
7]. Most approaches need a very dense
sampling of scale space and are therefore computationally expensive. Furthermore,
the connection with biological vision systems is not clear. We propose
here a single and simple neuronal mechanism for utilizing scale space:
the tracing and merging of contour sheets in scale space by neuronal oscillators.
Within this ansatz, neurons distributed over scale space at discrete points
![]() can participate in a common slow modulation of their mean firing rate,
if they code approximately the same intensity level. By this process, each
contour-sheet in scale space is can be marked with a specific "timecode".
These patches of synchronization are allowed to merge further, exhibiting
a common frequency, if -- in scale space -- no pronounced border
exists between them. As we will show, such a process is able to segment
and mark data into a few object chunks.
can participate in a common slow modulation of their mean firing rate,
if they code approximately the same intensity level. By this process, each
contour-sheet in scale space is can be marked with a specific "timecode".
These patches of synchronization are allowed to merge further, exhibiting
a common frequency, if -- in scale space -- no pronounced border
exists between them. As we will show, such a process is able to segment
and mark data into a few object chunks.
| Prev: Scale Space | ^ Up: Table of Contents ^ | Next: Neural Implementation |
Comments are welcome! © 1997 by Rolf Henkel - all rights reserved.