Research
Large-Scale Neural Recordings
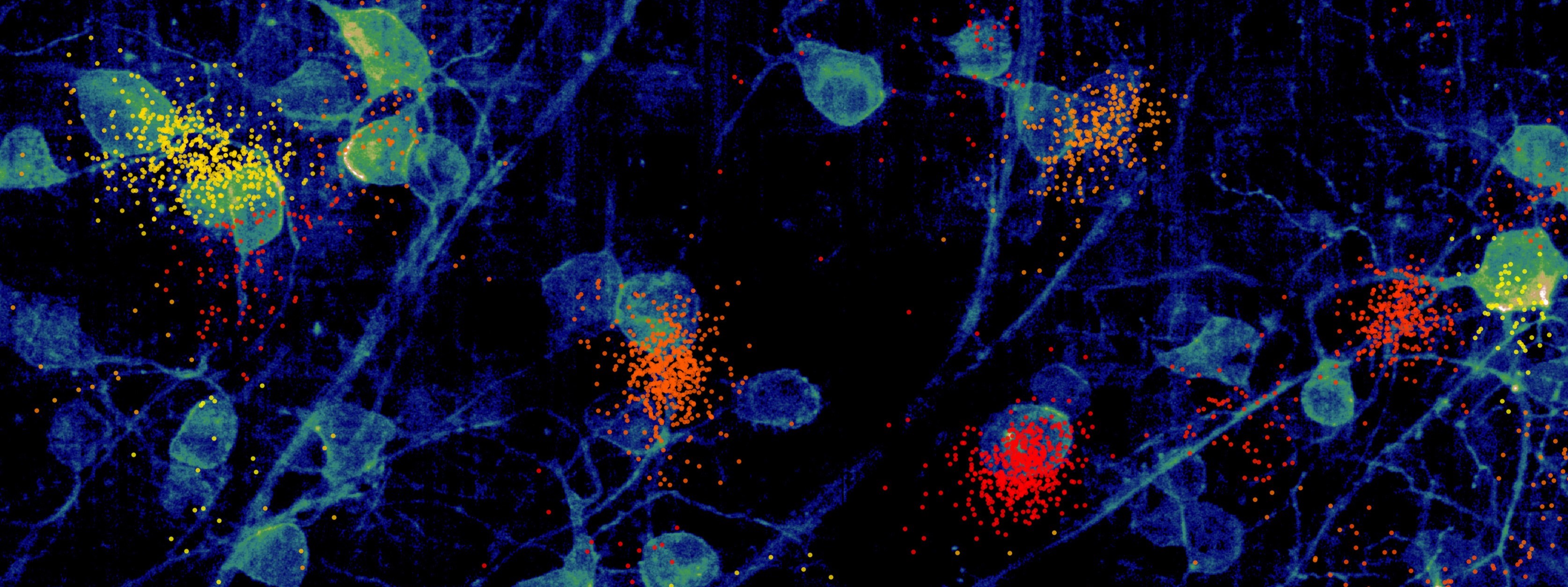
Next-generation, large-scale and dense neural recording systems now make it possible to precisely record the activity of 1000s neurons at the same time, a remarkable improvement over the handful of neurons most labs could at most record from only a decade ago. This technology gives us remarkable new opportunities to understand the inner workings of the brain, and as a result we are already witnessing a new focus on neural circuits and population dynamics.
Yet new technologies usually also bring new challenges. In the case of large-scale extracellular recordings, these are due to data volume and complexity. For instance, a Neuropixels 1 probe with 384 recording channels, now used in many labs, produces 23MB of raw data per second, so 82 GB/hour. A 3Brain Biocam with 4,096 channels yields 632 GB/s. Clearly fast algorithms are required to analyse such data, but a major problem is that the computational complexity does not scale linearly with the number of channels. The main challenge in this context is spike sorting: the assignment of detected events to single neurons.
As part of a community effort to tackle these problems, we develop tools for efficient, precise and reproducible analysis of large-scale neural recordings:
- The SpikeInterface project unifies existing and new spike sorting technologies into a single code base, for easy construction of analysis pipeleines and fully reproducible analysis.
- The Herding Spikes Project provides algorithms for neuronal spike detection and sorting, specifically optimised for high-density recording platforms with thousands of channels. It makes use of spatial spike localisation, exploiting the spatial signal spread across recording channels.
- As we found estimated spike locations to be an extremely useful low-dimensional feature, we developed a Bayesian approach to more reliably estimate them (compared to a naive centre-of-mass, which fails especially at the borders of recording probes). To make this scalable, we used amortised inference through VAE, which works very well even for gigantic data sets with 10s of millions of events: Scalable Spike Source Localization in Extracellular Recordings using Amortized Variational Inference
Modelling Neural Population Activity
Large-scale recordings are brilliant because they give us a glimpse of the network activity in much larger circuits that may just be sufficient to draw conclusions about its dynamics. This allows us to depart from the traditional approach of analysing recordings neuron by neuron, for instance by establishing which external covariates lead to a change in the activity of a neuron. Instead, we would like to make use of the fact that neural circuit dynamics are highly coordinated: neurons are not independently active, but they influence each other through their connections. This has important consequences that we can study with or even embed into statistical models. First, we expect the activity repertoire of a circuit to be limited, not all patterns of activity are possible because neurons excite or inhibit each other, albeit in complicated ways. Second, we expect that the temporal evolution of the activity is equally constrained: the connectivity of the network and the current state and the past history of the activity determine which states are accessible next.
These principles allow us to formulate principles and hypotheses for models of neural activity. One research direction is data-driven modelling, where we learn parametric models using neural data:
- We introduced Targeted Neural Dynamical Modeling as a new, weakly supervised method to discover behaviourally relevant latent dynamics in neural activity (Code).
- To align recordings from different sessions, we developed domain adaptation and self-supervised methods.
Some of these ideas and more are summarised in the following reviews:
- M.H. Hennig (2022). The sloppy relationship between neural circuit structure and function. Journal of Physiology. Early view.
- C. Hurwitz, N. Kudryashova, A. Onken, M.H. Hennig (2021). Building population models for large-scale neural recordings: opportunities and pitfalls. Current Opinion in Neurobiology, 70, 64-73.
Sloppiness in neuronal networks
Long term recordings from intact brains suggest that neurons are not stable, but subject to ongoing plasticity. This can be seen on different levels, for instance changes in spine sizes, or fluctuations in average activity levels. We observed the same phenomenon in high-density recordings from cell cultures, where individual neurons could change their activity and correlation with other neurons over the course of days.
This seems at odds with the basic postulate of neural homeostasis, which states that neurons are kept close to set-point activity levels. It also raises the question for stable function is maintained in presence of such changes.
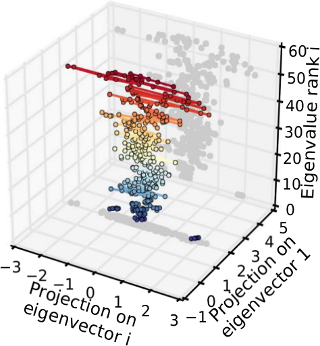
We found that the effects of single neurons on the activity generated by a whole network is highly anisotropic, with some ‘stiff’ neurons providing a stable backbone, around which fluctuations can occur in ‘sloppy’ neurons that have only a minor effect on the overall population activity. In terms of a parametric statistical model of population activity, this corresponds to a separation of parameter directions with high likelihood curvature, the stiff dimensions, and sloppy directions where the likelihood is essentially flat. Moreover, fluctuations over time occurred preferentially along sloppy directions, hence essential functional network properties were preserved over time.
This analysis was inspired by work in James Sethna’s group suggesting that sloppiness is a generic feature of complex, coupled non-linear systems.
D. Panas, H. Amin, A. Maccione, O. Muthmann, M. van Rossum, L. Berdondini, M.H. Hennig (2015). Sloppiness in spontaneously active neuronal networks. J Neurosci, 35(22): 8480-8492.
Retinal Development and Function
Retinal waves are a fascinating example of spontaneous network activity, which is found in developing neural circuits all over the brain. They consist of intrinsically generated propagating activity patterns, which disappear once the retinal circuits are ready for normal vision. Retinal waves have been found in many vertebrate species, and are thought to play an important role in guiding the neural development of the retina and the higher visual system. We study the regulation and developmental maturation of retinal waves, as well as the effects of their manipulation on retinal development. Using a combination of modelling and experiment, we found that the network producing early-stage cholinergic waves is exhibits dynamics close to a transition between local and global functional connectedness, and that this network state is under homeostatic control.
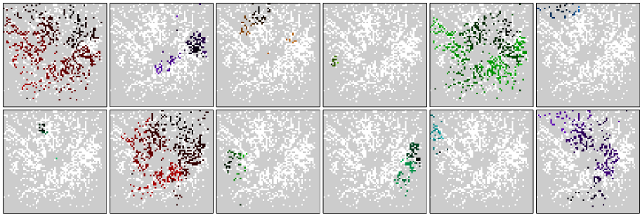

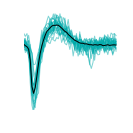
Twelve consecutive retinal waves recorded with the APS MEA in a P4 mouse retina, in presence of 6mM potassium in the solution. Waves travel from dark to light colours.